Contents |
General Remarks
HDF5 is a library for storing large numerical data sets. HDF stands for Hierarchical Data Format. It was designed for saving and retrieving data to/from structured large files. It also supports parallel access to files in HDF5 format, in particular within the MPI environment. HDF5 library can be used from C, C++ (with some limitations) and Fortran. Files can be saved optionally in text format, binary and compressed format (if zlib is available).
HDF5 is recommended by the developers of FFTW as a means to output data from different MPI processes.
Some popular numerical packages, like MATLAB, Mathematica, Octave and ROOT, already have native support for the HDF5 format. If you want to quickly experiment with HDF5 files, you can use those programs to see how it works.
For example, in Mathematica:
m = RandomInteger[255, {5, 5}]
Export[ "matrix.h5", m]
mLoad = Import["matrix.h5", {"Datasets", "/Dataset1"}]
will create a binary file called "matrix.h5" with the matrix data.
Install
The following are instructions to install HDF5 in different systems. Note that, as of HDF5 version 1.9 there is no way to use the MPI version and the C++ interfaces together. The C interface can be used from C++ anyway.
Ubuntu
HDF5 1.6.6 can be installed directly in Ubuntu 8.10 be doing:
sudo apt-get install libhdf5-serial-dev
This will install the C, C++ and Fortran versions of the library and development (header) files, but it will not include the MPI version.
The MPI version, (which will remove the previous serial version) can be installed by:
sudo apt-get install libhdf5-mpich-dev
where 'mpich' can be replaced by 'openmpi' or 'lam'. Unfortunately the C++ interface is not provided for this MPI version. The two version are incompatible.
Build and Installation from Sources
We will try to install the parallel version of HDF5 1.9 in our user space. (HDF5 1.8 --official release-- does not play well when compiling with gcc4.)
mk $HOME/usr
from a download location
mk $HOME/soft cd $HOME/soft wget ftp://ftp.hdfgroup.uiuc.edu/pub/outgoing/hdf5/snapshots/v19/hdf5-1.9.29.tar.gz tar -zxvf hdf5-1.9.29.tar.gz cd hdf5-1.9.29
then we can configure:
CC=mpicc ./configure --prefix=$HOME/usr --enable-parallel
Other options are described in ./configure --help. The option --enable-cxx can be specified but not together with --enable-parallel. For non-parallel version adding "CC=mpicc" (or equivalent) is not necessary. Check that your MPI compiler by doing, for example:
which mpicc
Then we can make and install
make make install
The compilation takes ~5 minutes and several warning messages will appear. Many header files will be installed in ~/usr/include and ~/usr/lib
~/usr/include/H5*.h (around 40 files) ~/usr/include/hdf5[|_hl].h ~/usr/lib/libhdf5[|_hl].[a|la]
The most important for us are hdf5.h and libhdf5.a. There are also some command line utilities to manage HDF5 files installed:
~/usr/bin/h5*
Among them, there is 'h5dump', which will be used in the next section.
Test Example
The source files contain examples on the usage of HDF5, including C++ examples. (See directories ./examples, ./hl/examples, ./c++/examples. and ./hl/c++/examples)
Simple introductory examples are also provided online, but they are outdated and are incompatible with this version of HDF5. This issue can be very confusing. It is better to use the examples contained in the distribution file and use the online documentation to read the details of the examples. In any case here I provide the sources I used for testing the installation h5_test.tar.gz. Use the example as follows:
wget http://micro.stanford.edu/mediawiki-1.11.0/images/H5_test.tar.gz -O h5_test.tar.gz tar -zxvf h5_test.tar.gz cd h5_test make test
Internally in the Makefile the compilation is performed by the command line:
mpicc -I${HOME}/usr/include h5_write.c -L${HOME}/usr/lib -lhdf5 -lm -lpng -o h5_write
A write and read program will be compiled. The write program will create a binary file named SDS.h5 with the data of a certain array, then this array will be loaded from the file by the read program and printed.
The SDS.h5 is in a compressed binary format, which means that it can not be read directly. However there exists a bunch of HDF5 utilities (external programs) that allows humans to see what is contained in the files:
$ ~/usr/bin/h5dump SDS.h5
HDF5 "SDS.h5" {
GROUP "/" {
DATASET "IntArray" {
DATATYPE H5T_STD_I32LE
DATASPACE SIMPLE { ( 5, 6 ) / ( 5, 6 ) }
DATA {
(0,0): 0, 1, 2, 3, 4, 5,
(1,0): 1, 2, 3, 4, 5, 6,
(2,0): 2, 3, 4, 5, 6, 7,
(3,0): 3, 4, 5, 6, 7, 8,
(4,0): 4, 5, 6, 7, 8, 9
}
}
}
}
For the moment this document is not a tutorial on HDF5 itself but it is only to document on its installation. However, we can already mention something about the structure of the file: For example, in the previous h5dump you can read 'GROUP "/"', this indicates that the dataset is at root level ('/') of the file. The HDF5 file can look pretty much like a filesystem, with directories, subdirectories and files/datasets.
Visualization Tools
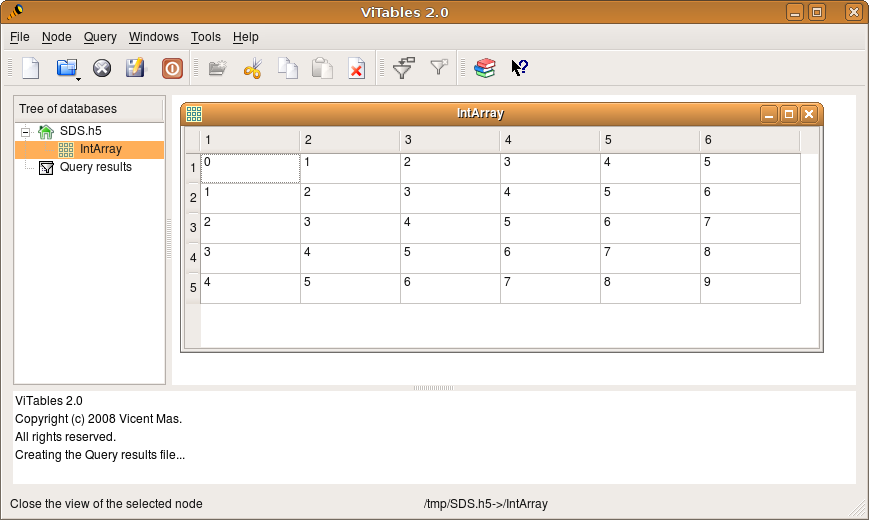
Note that last quoted code is not the real content of the file but just a human readable translation, that was accessed by means of one of the tools installed in ~/usr/bin. There are even graphical programs to visualize the contents of HDF5 file. The official one is HDFView (not tested here). An alternative that I tested is ViTables.
cd ~/soft wget http://prdownload.berlios.de/vitables/ViTables-2.0.tar.gz tar -zxvf ViTables-2.0.tar.gz cd ViTables-2.0 sudo apt-get python-tables python-qt4 sudo python setup.py install vitables
These programs that read HDF5 have the advantage that they do not load the data into memory. You can visualize the contents of a 4GB file without hanging the program.